Quantum Chemistry with OpenFermion
Set up molecular Hamiltonians in OpenFermion, apply Jordan-Wigner mapping to convert fermionic operators to Pauli strings, and build a VQE circuit in Cirq.
Circuit diagrams
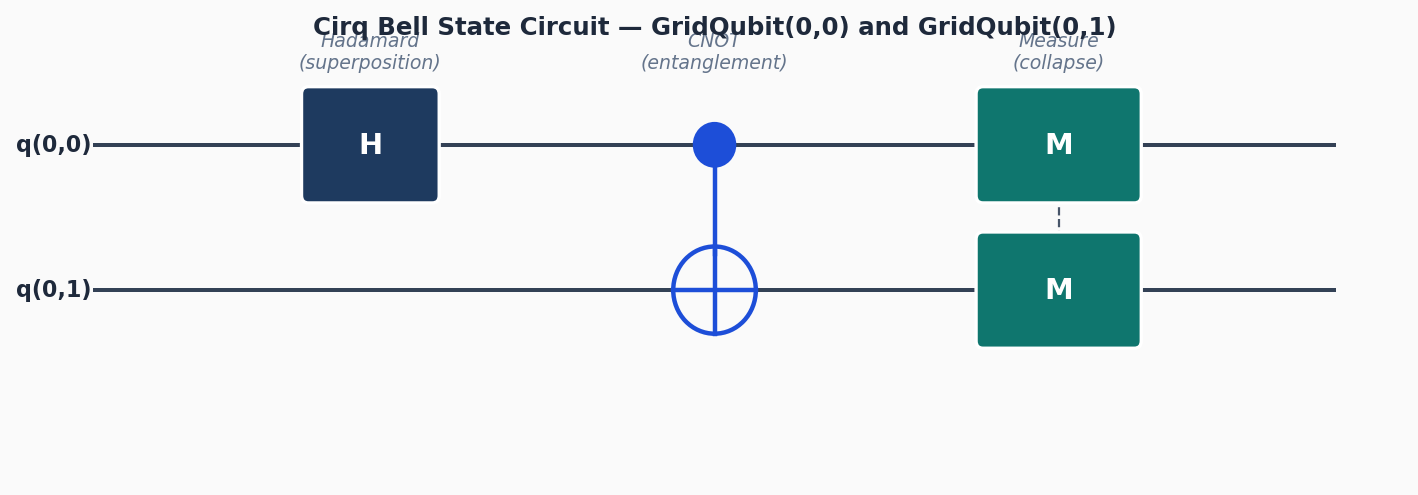
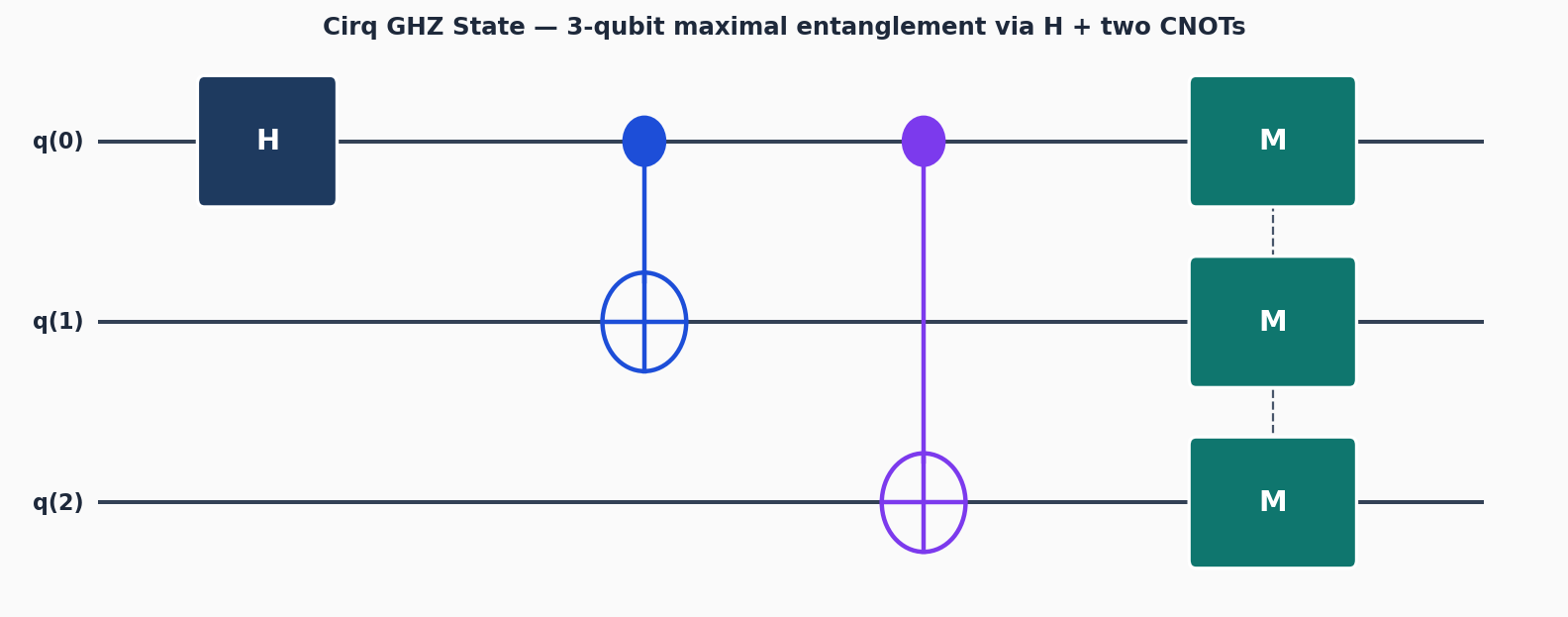
Simulating molecules is one of the most compelling applications for quantum computers. Classical Full Configuration Interaction (FCI) scales exponentially in the number of electrons, making it impractical beyond small systems. Quantum computers can, in principle, represent the electronic wavefunction efficiently. OpenFermion is the bridge: it lets you write fermionic Hamiltonians, transform them to qubit operators, and hand the result to a circuit framework like Cirq.
This tutorial walks through the complete pipeline, from second-quantized operators to VQE circuits, with runnable code at every step.
Installation
pip install openfermion openfermion-pyscf cirq pyscf
Why Second Quantization?
In first quantization, the electronic wavefunction is a function of all electron coordinates: Psi(r1, r2, …, rN). For N electrons, this function must be antisymmetric under exchange of any two electrons:
Psi(…, ri, …, rj, …) = -Psi(…, rj, …, ri, …)
This antisymmetry requirement (the Pauli exclusion principle) makes the bookkeeping difficult. For N electrons distributed among M spin orbitals, the Hilbert space has dimension C(M, N), the binomial coefficient “M choose N.” Representing the wavefunction as a linear combination of Slater determinants enforces the antisymmetry, but every integral involves determinants of overlap matrices.
Second quantization takes a different approach. Instead of tracking where each electron is, you track which orbitals are occupied. The state of the system is described by an occupation number vector:
|n_0, n_1, …, n_{M-1}>
where n_p is 0 or 1 (since electrons are fermions). Two operators act on this space:
- The creation operator a†_p places an electron in orbital p (if it is empty) and produces zero if orbital p is already occupied.
- The annihilation operator a_p removes an electron from orbital p (if one is present) and produces zero if orbital p is empty.
These operators satisfy the anticommutation relations:
{a_p, a†_q} = a_p a†_q + a†_q a_p = delta_pq
{a_p, a_q} = {a†_p, a†_q} = 0
These relations encode the Pauli exclusion principle algebraically. Any electronic Hamiltonian can be written in terms of these operators, and the formalism maps naturally to qubits because each spin orbital has two states (occupied or empty), just like a qubit has two states (|0> or |1>).
FermionOperator Basics
OpenFermion represents second-quantized operators with the FermionOperator class. The notation uses integers for orbital indices and ^ (caret) for creation operators.
from openfermion import FermionOperator, hermitian_conjugated, normal_ordered
# Creation and annihilation operators
a_dag_0 = FermionOperator('0^', 1.0) # a†_0: create in orbital 0
a_1 = FermionOperator('1', 1.0) # a_1: destroy in orbital 1
# Hopping term: move electron from orbital 1 to orbital 0
hop = FermionOperator('0^ 1', 1.0)
# Number operator: n_0 = a†_0 a_0 (counts electrons in orbital 0)
n_0 = FermionOperator('0^ 0', 1.0)
# Normal ordering pushes all creation operators to the left,
# picking up signs from anticommutation.
print(normal_ordered(a_1 * a_dag_0))
# -1.0 [0^ 1]
The output -1.0 [0^ 1] shows anticommutation in action: swapping a_1 and a†_0 introduces a minus sign (since p != q, the anticommutator is zero, so a_1 a†_0 = -a†_0 a_1).
FermionOperator Arithmetic
FermionOperators support addition, subtraction, scalar multiplication, operator multiplication, and Hermitian conjugation.
from openfermion import FermionOperator, hermitian_conjugated, normal_ordered
# Hermitian conjugate of a hopping term
hop = FermionOperator('0^ 1', 1.0)
hop_dag = hermitian_conjugated(hop)
print(f"hop = {hop}") # 1.0 [0^ 1]
print(f"hop_dag = {hop_dag}") # 1.0 [1^ 0]
# A Hermitian hopping term (required for physical Hamiltonians)
hop_hermitian = hop + hop_dag
print(f"hop + h.c. = {hop_hermitian}")
# 1.0 [0^ 1] + 1.0 [1^ 0]
# Scalar multiplication
print(f"2 * hop = {2.0 * hop}") # 2.0 [0^ 1]
# Operator multiplication (composition)
product = hop * hop_dag
print(f"hop * hop_dag = {product}")
print(f"normal ordered = {normal_ordered(product)}")
# The normal-ordered form reveals the physics:
# a†_0 a_1 a†_1 a_0 = a†_0 (delta_11 - a†_1 a_1) a_0
# = a†_0 a_0 - a†_0 a†_1 a_1 a_0
# Number operator for multiple orbitals
total_number = sum(FermionOperator(f'{i}^ {i}', 1.0) for i in range(4))
print(f"Total number operator (4 orbitals): {total_number}")
# Count the terms in an operator
print(f"Number of terms in hop_hermitian: {len(hop_hermitian.terms)}")
Normal Ordering Matters
Normal ordering places all creation operators to the left of all annihilation operators within each term, applying anticommutation relations as needed. This produces a canonical form that makes two operators comparable.
from openfermion import FermionOperator, normal_ordered
# These represent the same physical operator but look different
op_a = FermionOperator('0^ 1^ 1 0', 1.0) # a†_0 a†_1 a_1 a_0
op_b = FermionOperator('1^ 0^ 0 1', 1.0) # a†_1 a†_0 a_0 a_1
# Direct comparison fails
print(f"Before normal ordering, equal: {op_a == op_b}") # False
# After normal ordering, both reduce to the same form
print(f"After normal ordering, equal: "
f"{normal_ordered(op_a) == normal_ordered(op_b)}") # True
Always call normal_ordered() before comparing FermionOperators for equality. Without it, two expressions for the same operator may appear different.
Jordan-Wigner Mapping
A quantum computer operates on qubits, not fermions. The Jordan-Wigner (JW) transformation maps fermionic operators to qubit (Pauli) operators, preserving the anticommutation algebra.
The mapping assigns one qubit per spin orbital. Orbital occupation maps directly to qubit state: occupied (1 electron) becomes |1>, empty (0 electrons) becomes |0>.
from openfermion import jordan_wigner, count_qubits, FermionOperator
# Hermitian hopping term
fermion_op = FermionOperator('0^ 1', 1.0) + FermionOperator('1^ 0', 1.0)
qubit_op = jordan_wigner(fermion_op)
print(qubit_op)
# (0.5+0j) [X0 X1] +
# (0.5+0j) [Y0 Y1]
print(f"Qubits required: {count_qubits(qubit_op)}") # 2
Each term in the resulting QubitOperator is a weighted Pauli string that can be measured on a quantum computer.
The Z String Explained
The key subtlety in Jordan-Wigner is the “Z string.” Fermionic operators anticommute, but qubit operators on different qubits commute. To encode the fermionic sign, JW inserts Z operators on all qubits between 0 and p-1 when mapping a†_p.
The rule is:
- a†p maps to (1/2)(X_p - iY_p) tensor Z{p-1} tensor Z_{p-2} tensor … tensor Z_0
- a_p maps to (1/2)(X_p + iY_p) tensor Z_{p-1} tensor Z_{p-2} tensor … tensor Z_0
The Z string counts the parity of occupied orbitals below site p. Since Z|0> = |0> and Z|1> = -|1>, each occupied orbital below p contributes a factor of -1, reproducing the fermionic sign.
You can see this directly by mapping higher-indexed operators:
from openfermion import FermionOperator, jordan_wigner
# Map a†_0: no Z string needed (no qubits below index 0)
print("a†_0 =", jordan_wigner(FermionOperator('0^', 1.0)))
# (0.5+0j) [X0] + (-0.5j) [Y0]
# Map a†_1: Z string on qubit 0
print("a†_1 =", jordan_wigner(FermionOperator('1^', 1.0)))
# (0.5+0j) [Z0 X1] + (-0.5j) [Z0 Y1]
# Map a†_2: Z string on qubits 0 and 1
print("a†_2 =", jordan_wigner(FermionOperator('2^', 1.0)))
# (0.5+0j) [Z0 Z1 X2] + (-0.5j) [Z0 Z1 Y2]
Each additional orbital index adds one more Z to the string. For an operator on orbital p, the JW-mapped term has Pauli weight p+1 (one X or Y on qubit p, plus p Z operators). This O(n) scaling in Pauli weight is the main drawback of the Jordan-Wigner transform: terms acting on high-indexed orbitals produce long Pauli strings that require many two-qubit gates to implement.
Bravyi-Kitaev Alternative
The Bravyi-Kitaev (BK) transform encodes occupation numbers in a binary tree structure. Instead of storing the occupation of each orbital directly, BK stores partial parity sums. This reduces the average Pauli weight from O(n) to O(log n).
For small systems the difference is modest, but it matters for fault-tolerant resource estimates on larger molecules where circuit depth is the bottleneck.
from openfermion import FermionOperator, jordan_wigner, bravyi_kitaev
fermion_op = FermionOperator('0^ 1', 1.0) + FermionOperator('1^ 0', 1.0)
qubit_op_jw = jordan_wigner(fermion_op)
qubit_op_bk = bravyi_kitaev(fermion_op)
print(f"JW: {qubit_op_jw}")
print(f"BK: {qubit_op_bk}")
# Compare average Pauli weights
def avg_weight(op):
weights = [len(t) for t in op.terms if t]
return sum(weights) / len(weights) if weights else 0
print(f"JW avg Pauli weight: {avg_weight(qubit_op_jw):.2f}")
print(f"BK avg Pauli weight: {avg_weight(qubit_op_bk):.2f}")
For a two-orbital example the weights are similar. The advantage appears at larger orbital counts where JW Z strings grow linearly while BK strings grow logarithmically.
QubitOperator Arithmetic
After transforming to qubit operators, you frequently need to manipulate them: add operators, compute commutators, extract individual Pauli terms, or convert to matrices.
from openfermion import (
QubitOperator, jordan_wigner, FermionOperator,
commutator, get_sparse_operator, count_qubits
)
import numpy as np
# Build two qubit operators directly
op_a = QubitOperator('X0 X1', 0.5)
op_b = QubitOperator('Z0', -0.3) + QubitOperator('Z1', -0.3)
# Addition
total = op_a + op_b
print(f"Sum: {total}")
# Multiplication (operator composition)
product = op_a * op_b
print(f"Product: {product}")
# Commutator [A, B] = AB - BA
comm = commutator(op_a, op_b)
print(f"Commutator: {comm}")
# Iterate over Pauli terms with their coefficients
fermion_op = FermionOperator('0^ 1', 1.0) + FermionOperator('1^ 0', 1.0)
qubit_op = jordan_wigner(fermion_op)
print("\nPauli terms in JW-mapped hopping operator:")
for term, coeff in qubit_op.terms.items():
pauli_str = ' '.join(f'{p}{i}' for i, p in term) if term else 'I'
print(f" {coeff:+.4f} * {pauli_str}")
# Convert to sparse matrix (scipy.sparse)
sparse_matrix = get_sparse_operator(qubit_op)
print(f"\nSparse matrix shape: {sparse_matrix.shape}")
print(f"Nonzero entries: {sparse_matrix.nnz}")
# For small systems, convert to dense and diagonalize
dense_matrix = sparse_matrix.toarray()
eigenvalues = np.linalg.eigvalsh(dense_matrix)
print(f"Eigenvalues: {eigenvalues}")
Note the string format difference: QubitOperator('X0 Y1') uses Pauli labels (X, Y, Z) with qubit indices, while FermionOperator('0^ 1') uses orbital indices with ^ for creation. Mixing these formats is a common source of bugs.
OpenFermion-PySCF Integration
For real molecules, you need one-electron and two-electron integrals computed from an atomic orbital basis set. The openfermionpyscf package provides the run_pyscf function, which handles this automatically.
What run_pyscf Does
Internally, run_pyscf performs these steps:
- Runs a Hartree-Fock (HF) calculation to find molecular orbitals
- Computes one-electron integrals h_pq (kinetic energy + nuclear attraction) in the molecular orbital basis
- Computes two-electron integrals g_pqrs (electron-electron repulsion) in the molecular orbital basis
- Optionally runs CCSD (coupled-cluster) and FCI for comparison energies
The output is a MolecularData object containing all computed quantities.
MolecularData, InteractionOperator, and FermionOperator
Three classes form a pipeline from raw chemistry data to qubit-ready operators:
- MolecularData stores molecular geometry, basis set, and computed integrals.
- InteractionOperator stores the Hamiltonian coefficients in a structured format: a constant (nuclear repulsion), one-body tensor h_pq, and two-body tensor h_pqrs.
- FermionOperator is the general second-quantized operator, ready for JW or BK transformation.
# Requires: pyscf
from openfermion import (
MolecularData, jordan_wigner, get_sparse_operator,
get_fermion_operator, count_qubits
)
from openfermionpyscf import run_pyscf
import numpy as np
# Define molecular geometry (coordinates in angstrom)
geometry = [('H', (0., 0., 0.)), ('H', (0., 0., 0.7414))]
molecule = MolecularData(geometry, basis='sto-3g', multiplicity=1, charge=0)
molecule = run_pyscf(molecule, run_fci=True)
print(f"Number of spatial orbitals: {molecule.n_orbitals}")
print(f"Number of spin orbitals (qubits): {molecule.n_qubits}")
print(f"Number of electrons: {molecule.n_electrons}")
print(f"HF energy: {molecule.hf_energy:.6f} Ha")
print(f"FCI energy: {molecule.fci_energy:.6f} Ha")
Accessing Molecular Integrals
The one-body integrals h_pq represent single-electron terms: kinetic energy and nuclear attraction. The two-body integrals g_pqrs represent electron-electron repulsion. Both are indexed by spatial orbitals.
# Requires: pyscf (continued from above)
# One-body integrals: h_pq (spatial orbital basis)
# These encode kinetic energy and electron-nuclear attraction
print(f"One-body integrals shape: {molecule.one_body_integrals.shape}")
print(f"One-body integrals (h_pq):\n{molecule.one_body_integrals}")
# For H2/STO-3G: 2x2 matrix (2 spatial orbitals)
# h_00 ~ -1.25 Ha (bonding orbital, lower energy)
# h_11 ~ -0.48 Ha (antibonding orbital, higher energy)
# Two-body integrals: g_pqrs (spatial orbital basis)
# These encode electron-electron Coulomb repulsion
print(f"\nTwo-body integrals shape: {molecule.two_body_integrals.shape}")
From Integrals to Qubit Hamiltonian
The get_molecular_hamiltonian() method returns an InteractionOperator that combines the nuclear repulsion constant, one-body integrals, and two-body integrals. The InteractionOperator uses spin-orbital indexing (even indices = alpha/spin-up, odd indices = beta/spin-down).
# Requires: pyscf (continued from above)
# Step 1: MolecularData -> InteractionOperator
hamiltonian = molecule.get_molecular_hamiltonian()
print(f"Nuclear repulsion (constant): {hamiltonian.constant:.6f} Ha")
print(f"One-body tensor shape: {hamiltonian.one_body_tensor.shape}")
print(f"Two-body tensor shape: {hamiltonian.two_body_tensor.shape}")
# The one-body tensor is in spin-orbital basis (4x4 for H2/STO-3G)
# Even indices (0, 2) are alpha spin; odd indices (1, 3) are beta spin
print(f"\nOne-body tensor (spin-orbital):\n{hamiltonian.one_body_tensor}")
# Step 2: InteractionOperator -> QubitOperator via Jordan-Wigner
qubit_hamiltonian = jordan_wigner(hamiltonian)
print(f"\nQubit Hamiltonian has {len(qubit_hamiltonian.terms)} Pauli terms")
print(f"Qubits required: {count_qubits(qubit_hamiltonian)}")
# Step 3: Convert to sparse matrix for classical diagonalization
H_sparse = get_sparse_operator(qubit_hamiltonian)
H_matrix = H_sparse.toarray()
# Verify: ground state eigenvalue matches FCI
eigenvalues = np.linalg.eigvalsh(H_matrix)
print(f"\nGround state from diag: {eigenvalues[0]:.6f} Ha")
print(f"FCI energy: {molecule.fci_energy:.6f} Ha")
You can also go from InteractionOperator to FermionOperator using get_fermion_operator():
# Requires: pyscf (continued from above)
fermion_hamiltonian = get_fermion_operator(hamiltonian)
print(f"FermionOperator has {len(fermion_hamiltonian.terms)} terms")
Symmetry Reduction with SCBK
H2 in STO-3G has 4 spin orbitals, requiring 4 qubits under Jordan-Wigner. But the molecular Hamiltonian conserves both total electron number and total spin. The symmetry-conserving Bravyi-Kitaev (SCBK) transformation exploits these two symmetries to remove 2 qubits, leaving only 2 qubits for H2.
# Requires: pyscf (continued from above)
from openfermion import symmetry_conserving_bravyi_kitaev
# Convert InteractionOperator to FermionOperator first
fermion_h = get_fermion_operator(molecule.get_molecular_hamiltonian())
# SCBK: specify total spin orbitals and total electrons
scbk_hamiltonian = symmetry_conserving_bravyi_kitaev(
fermion_h,
active_orbitals=4, # number of spin orbitals
active_fermions=2 # number of electrons
)
print(f"JW qubits: {count_qubits(qubit_hamiltonian)}")
print(f"SCBK qubits: {count_qubits(scbk_hamiltonian)}")
print(f"JW Pauli terms: {len(qubit_hamiltonian.terms)}")
print(f"SCBK Pauli terms: {len(scbk_hamiltonian.terms)}")
# Print the reduced Hamiltonian
print("\nSCBK Hamiltonian:")
for term, coeff in scbk_hamiltonian.terms.items():
if abs(coeff) > 1e-10:
pauli_str = ' '.join(f'{p}{i}' for i, p in term) if term else 'I'
print(f" {coeff:+.6f} * {pauli_str}")
# Verify ground state energy is preserved
H_scbk = get_sparse_operator(scbk_hamiltonian).toarray()
eigvals = np.linalg.eigvalsh(H_scbk)
print(f"\nSCBK ground state: {eigvals[0]:.6f} Ha")
print(f"FCI energy: {molecule.fci_energy:.6f} Ha")
For H2, SCBK reduces the Hamiltonian from 15 Pauli terms on 4 qubits to 5 Pauli terms on 2 qubits. The ground state energy is exactly preserved. This reduction becomes even more valuable for larger molecules where every qubit saved is significant.
Hamiltonian Grouping for VQE Measurements
In a VQE experiment on real hardware, you measure the expectation value of each Pauli term in the Hamiltonian. Measuring each term individually requires many circuit executions. However, Pauli terms that are diagonal in the same tensor product basis can be measured simultaneously with a single circuit.
OpenFermion’s group_into_tensor_product_basis_sets partitions the Hamiltonian into groups of terms that share a common measurement basis.
# Requires: pyscf (continued from above)
from openfermion import group_into_tensor_product_basis_sets
# Group the 4-qubit JW Hamiltonian
groups = group_into_tensor_product_basis_sets(qubit_hamiltonian)
print(f"Total Pauli terms: {len(qubit_hamiltonian.terms)}")
print(f"Measurement groups needed: {len(groups)}")
print()
for basis, sub_op in groups.items():
# The basis key describes the single-qubit rotations needed
basis_str = ', '.join(f'{p}{i}' for i, p in basis)
print(f"Basis ({basis_str}): {len(sub_op.terms)} terms")
For H2 with 15 Pauli terms, grouping typically yields 5 measurement bases. On hardware, this means 5 distinct circuits (each with different pre-measurement rotations) instead of 15. For larger molecules with hundreds of terms, grouping can reduce the measurement overhead by an order of magnitude.
Run VQE with SciPy (Full 4-Qubit)
With the Hamiltonian as a matrix, VQE minimizes the expectation value over UCCSD amplitudes using scipy.optimize.minimize. This approach uses exact statevector simulation (no sampling noise) and works well for small systems.
Requires:
H_matrixfrom the PySCF block above.
# Requires: pyscf
import numpy as np
from scipy.optimize import minimize
from scipy.linalg import expm
from openfermion import FermionOperator, jordan_wigner, get_sparse_operator
n_qubits = 4
n_electrons = 2
# Single excitations: (occupied, virtual) orbital pairs
# Occupied spin orbitals: 0, 1; Virtual: 2, 3
singles = [(0, 2), (0, 3), (1, 2), (1, 3)]
# Double excitations: (occ1, occ2, virt1, virt2)
doubles = [(0, 1, 2, 3)]
def build_uccsd_generator(t1, t2):
"""Build the anti-Hermitian UCCSD cluster operator T - T†."""
gen = FermionOperator()
for (i, a), theta in zip(singles, t1):
gen += FermionOperator(f'{a}^ {i}', theta)
gen -= FermionOperator(f'{i}^ {a}', theta)
for (i, j, a, b), theta in zip(doubles, t2):
gen += FermionOperator(f'{a}^ {b}^ {j} {i}', theta)
gen -= FermionOperator(f'{i}^ {j}^ {b} {a}', theta)
return gen
# Hartree-Fock reference state: orbitals 0 and 1 occupied
# In JW encoding, this is |0011> = |3> (little-endian: qubit 0 is rightmost)
psi_hf = np.zeros(2 ** n_qubits, dtype=complex)
psi_hf[3] = 1.0
def energy(params):
"""Compute <psi|H|psi> where |psi> = exp(i * JW(T-T†)) |HF>."""
t1 = params[:len(singles)]
t2 = params[len(singles):]
gen = build_uccsd_generator(t1, t2)
jw_gen = get_sparse_operator(jordan_wigner(gen), n_qubits=n_qubits).toarray()
psi = expm(1j * jw_gen) @ psi_hf
return np.real(psi.conj() @ H_matrix @ psi)
x0 = np.zeros(len(singles) + len(doubles))
result = minimize(energy, x0, method='BFGS', options={'gtol': 1e-6})
print(f"VQE energy: {result.fun:.6f} Ha")
print(f"Parameters: {len(x0)} (4 singles + 1 double)")
Compare to Exact FCI
# Requires: pyscf
fci_energy = np.linalg.eigvalsh(H_matrix)[0]
print(f"VQE energy: {result.fun:.6f} Ha")
print(f"FCI energy: {fci_energy:.6f} Ha")
print(f"Error: {abs(result.fun - fci_energy) * 1000:.3f} mHa")
# VQE energy: -1.137270 Ha
# FCI energy: -1.137270 Ha
# Error: 0.000 mHa
For H2, UCCSD is essentially exact: the singles and doubles amplitudes capture all electron correlation in a two-electron system. Chemical accuracy is 1.6 mHa (1 kcal/mol); the VQE result is well inside that threshold.
VQE with a Cirq Circuit (2-Qubit H2)
The matrix-based VQE above is useful for prototyping, but a real quantum computer executes circuits. Here we build an actual Cirq circuit for the 2-qubit SCBK-reduced H2 Hamiltonian.
After SCBK reduction, the H2 Hamiltonian has only 5 terms on 2 qubits. The UCCSD ansatz in this reduced space simplifies to a single parameter: a rotation that mixes the reference state |11> with the doubly-excited state |00>.
# Requires: pyscf
import cirq
import numpy as np
from scipy.optimize import minimize
from openfermion import (
MolecularData, get_sparse_operator, get_fermion_operator,
symmetry_conserving_bravyi_kitaev
)
from openfermionpyscf import run_pyscf
# Build 2-qubit Hamiltonian via SCBK
geometry = [('H', (0., 0., 0.)), ('H', (0., 0., 0.7414))]
molecule = MolecularData(geometry, basis='sto-3g', multiplicity=1, charge=0)
molecule = run_pyscf(molecule, run_fci=True)
fermion_h = get_fermion_operator(molecule.get_molecular_hamiltonian())
scbk_h = symmetry_conserving_bravyi_kitaev(
fermion_h, active_orbitals=4, active_fermions=2
)
H_matrix = get_sparse_operator(scbk_h).toarray()
# Define the Cirq circuit
q0, q1 = cirq.LineQubit.range(2)
def build_circuit(theta):
circuit = cirq.Circuit()
# Prepare |11>: the Hartree-Fock reference in SCBK encoding
circuit.append(cirq.X(q0))
circuit.append(cirq.X(q1))
# UCCSD ansatz: mix |11> with |00> using a CNOT-Ry-CNOT sequence
# The first CNOT maps |11> to |10>, putting the "excitation" on q0
circuit.append(cirq.CNOT(q0, q1))
# Ry rotation on q0 creates superposition of |0> and |1>
circuit.append(cirq.ry(theta).on(q0))
# Second CNOT entangles back: maps to superposition of |11> and |00>
circuit.append(cirq.CNOT(q0, q1))
return circuit
# Print the circuit at a sample angle
print("Circuit structure:")
print(build_circuit(0.2))
# VQE optimization loop
simulator = cirq.Simulator()
def energy_from_cirq(params):
theta = params[0]
circuit = build_circuit(theta)
result = simulator.simulate(circuit)
psi = result.final_state_vector
return np.real(psi.conj() @ H_matrix @ psi)
result = minimize(energy_from_cirq, x0=[0.0], method='COBYLA')
print(f"\nCirq VQE energy: {result.fun:.6f} Ha")
print(f"FCI energy: {molecule.fci_energy:.6f} Ha")
print(f"Error: {abs(result.fun - molecule.fci_energy) * 1000:.3f} mHa")
print(f"Optimal theta: {result.x[0]:.6f} rad")
This circuit has depth 5 and uses only 1 CNOT gate layer (the minimum for creating entanglement). On real hardware, you would replace the exact statevector simulation with repeated measurements, estimating each Pauli term’s expectation value from shot statistics.
Potential Energy Surface Scan
A potential energy surface (PES) shows how the total energy varies with molecular geometry. For H2, the relevant coordinate is the bond length. Scanning from 0.3 to 3.0 angstrom reveals the equilibrium geometry (energy minimum) and the dissociation limit (where the bond breaks).
# Requires: pyscf
from openfermion import MolecularData, jordan_wigner, get_sparse_operator
from openfermionpyscf import run_pyscf
import numpy as np
distances = np.arange(0.3, 3.1, 0.1)
hf_energies = []
fci_energies = []
for d in distances:
geometry = [('H', (0., 0., 0.)), ('H', (0., 0., float(d)))]
mol = MolecularData(geometry, basis='sto-3g', multiplicity=1, charge=0)
mol = run_pyscf(mol, run_fci=True)
hf_energies.append(mol.hf_energy)
fci_energies.append(mol.fci_energy)
# Find equilibrium geometry
min_idx = np.argmin(fci_energies)
print(f"Equilibrium bond length: {distances[min_idx]:.2f} angstrom")
print(f"Equilibrium FCI energy: {fci_energies[min_idx]:.6f} Ha")
print(f"Dissociation limit: {fci_energies[-1]:.6f} Ha")
# Print table for plotting
print("\n Bond (A) | HF (Ha) | FCI (Ha) | Correlation (mHa)")
print("-" * 62)
for d, hf, fci in zip(distances, hf_energies, fci_energies):
corr = (fci - hf) * 1000 # correlation energy in mHa
print(f" {d:5.2f} | {hf:11.6f} | {fci:11.6f} | {corr:10.3f}")
Two features stand out in the PES:
- The equilibrium bond length (~0.74 angstrom, FCI energy ~-1.137 Ha) matches experimental data for H2 in this basis set.
- At large bond distances, the HF and FCI curves diverge sharply. HF fails to describe bond breaking because it uses a single determinant. FCI captures the multi-reference character needed for dissociation. This is precisely the regime where quantum computers offer an advantage.
To plot the PES, pass the arrays to matplotlib:
# Optional: requires matplotlib
import matplotlib.pyplot as plt
plt.figure(figsize=(8, 5))
plt.plot(distances, hf_energies, 'b--', label='Hartree-Fock')
plt.plot(distances, fci_energies, 'r-', label='FCI')
plt.xlabel('Bond length (angstrom)')
plt.ylabel('Energy (Hartree)')
plt.title('H2 Potential Energy Surface (STO-3G)')
plt.legend()
plt.grid(True, alpha=0.3)
plt.tight_layout()
plt.savefig('h2_pes.png', dpi=150)
plt.show()
Larger Molecule Example: LiH
Moving beyond H2, lithium hydride (LiH) provides a more realistic test case. In the STO-3G basis, LiH has 6 spatial orbitals (12 spin orbitals), requiring 12 qubits under Jordan-Wigner.
# Requires: pyscf
from openfermion import MolecularData, jordan_wigner, count_qubits, get_fermion_operator
from openfermionpyscf import run_pyscf
geometry = [('Li', (0., 0., 0.)), ('H', (0., 0., 1.6))]
molecule = MolecularData(geometry, basis='sto-3g', multiplicity=1, charge=0)
molecule = run_pyscf(molecule, run_fci=True)
print(f"Spatial orbitals: {molecule.n_orbitals}")
print(f"Spin orbitals: {molecule.n_qubits}")
print(f"Electrons: {molecule.n_electrons}")
print(f"HF energy: {molecule.hf_energy:.6f} Ha")
print(f"FCI energy: {molecule.fci_energy:.6f} Ha")
print(f"Correlation: {(molecule.fci_energy - molecule.hf_energy)*1000:.3f} mHa")
12 qubits is already challenging for VQE on current hardware. The full UCCSD ansatz for 4 electrons in 12 spin orbitals has dozens of parameters, and each parameter corresponds to a circuit block with multiple CNOT gates.
Active Space Reduction
The lithium 1s core orbital (spatial orbital 0) is tightly bound and barely participates in bonding. Freezing it reduces the problem size without sacrificing much accuracy.
# Requires: pyscf (continued from above)
# Freeze spatial orbital 0 (Li 1s core, both alpha and beta)
# Active space: spatial orbitals 1, 2, 3, 4, 5
# This gives 10 spin orbitals with 2 active electrons
hamiltonian_full = molecule.get_molecular_hamiltonian()
hamiltonian_as = molecule.get_molecular_hamiltonian(
occupied_indices=[0], # freeze orbital 0
active_indices=[1, 2, 3, 4, 5] # keep orbitals 1-5 active
)
jw_full = jordan_wigner(hamiltonian_full)
jw_as = jordan_wigner(hamiltonian_as)
print(f"Full space: {count_qubits(jw_full)} qubits, "
f"{len(jw_full.terms)} Pauli terms")
print(f"Active space: {count_qubits(jw_as)} qubits, "
f"{len(jw_as.terms)} Pauli terms")
# Verify the active space Hamiltonian still gives a reasonable energy
import numpy as np
from openfermion import get_sparse_operator
H_as = get_sparse_operator(jw_as).toarray()
eigvals = np.linalg.eigvalsh(H_as)
print(f"\nActive space ground state: {eigvals[0]:.6f} Ha")
print(f"Full FCI energy: {molecule.fci_energy:.6f} Ha")
print(f"Active space error: "
f"{abs(eigvals[0] - molecule.fci_energy)*1000:.3f} mHa")
The active space approximation introduces a small error (typically < 1 mHa for LiH) while cutting the qubit count significantly. For VQE on near-term hardware, this trade-off is essential.
UCCSD Parameters for LiH Active Space
In the active space with 2 electrons and 10 spin orbitals, the UCCSD ansatz has:
- Singles: each occupied spin orbital can excite to each virtual spin orbital. With 2 occupied and 8 virtual spin orbitals (respecting spin symmetry: 1 alpha occupied to 4 alpha virtual, 1 beta occupied to 4 beta virtual), there are 8 single excitation parameters.
- Doubles: pairs of occupied spin orbitals excite to pairs of virtual spin orbitals. For alpha-beta doubles (the dominant contribution in a singlet state), there are 4 x 4 = 16 combinations, but spin symmetry and spatial symmetry reduce this. The total is typically around 8-12 independent double excitation parameters.
Each parameter translates to a block of gates in the quantum circuit. More parameters means deeper circuits and more measurements, which is why active space selection is critical for practical VQE.
Common Mistakes
1. Mixing QubitOperator and FermionOperator string formats
from openfermion import FermionOperator, QubitOperator
# Correct: FermionOperator uses orbital indices with ^ for creation
fermion = FermionOperator('0^ 1', 1.0)
# Correct: QubitOperator uses Pauli labels with qubit indices
qubit = QubitOperator('X0 Y1', 0.5)
# WRONG: do not use Pauli labels in FermionOperator
# FermionOperator('X0 Y1', 0.5) # raises an error
# WRONG: do not use ^ notation in QubitOperator
# QubitOperator('0^ 1', 1.0) # raises an error
2. Forgetting normal ordering before comparison
from openfermion import FermionOperator, normal_ordered
op1 = FermionOperator('0^ 1^ 1 0', 1.0)
op2 = FermionOperator('1^ 0^ 0 1', 1.0)
# Without normal ordering, these look different
assert op1 != op2
# After normal ordering, they are equal
assert normal_ordered(op1) == normal_ordered(op2)
3. Memory limits with get_sparse_operator
get_sparse_operator returns a scipy sparse matrix of size 2^n x 2^n. For n = 12 qubits, the dense matrix uses 2^12 x 2^12 x 16 bytes = 256 MB. For n = 20, it would need 16 TB. Use .toarray() only for small systems (up to ~14 qubits). For larger systems, use iterative eigensolvers like scipy.sparse.linalg.eigsh that work directly with the sparse format:
from scipy.sparse.linalg import eigsh
from openfermion import get_sparse_operator
# For large systems, use eigsh instead of converting to dense
# H_sparse = get_sparse_operator(large_qubit_hamiltonian)
# eigenvalue, eigenvector = eigsh(H_sparse, k=1, which='SA')
4. Unit confusion between PySCF and OpenFermion
PySCF expects geometry input in angstrom by default, and MolecularData follows the same convention. However, some quantum chemistry codes use Bohr (1 Bohr = 0.529177 angstrom). Always verify the unit convention when importing geometries from external sources.
5. Qubit ordering in circuits
Qiskit uses little-endian qubit ordering (qubit 0 is the least significant bit), while Cirq uses LineQubit indices that map to tensor product ordering (qubit 0 is the most significant bit in the state vector by default). When mapping OpenFermion’s QubitOperator to circuit measurement bases, verify that your qubit indexing convention matches the operator’s convention. A mismatch silently produces wrong expectation values.
What to Try Next
- Apply SCBK reduction to LiH and run VQE on the reduced qubit Hamiltonian
- Compare BK vs. Jordan-Wigner circuit depth using
openfermion-cirq’s circuit generators - Implement error mitigation on the Cirq VQE circuit (see the Mitiq tutorial)
- Use
scipy.sparse.linalg.eigshto compute ground states for systems too large for dense diagonalization - See the OpenFermion Reference for the full operator and transformation API
Was this tutorial helpful?