VQE with Cirq and OpenFermion
Build a complete VQE implementation for H2 using Cirq's parametric gates, OpenFermion for the qubit Hamiltonian, scipy for classical optimization, and the parameter shift rule for exact gradients.
Circuit diagrams
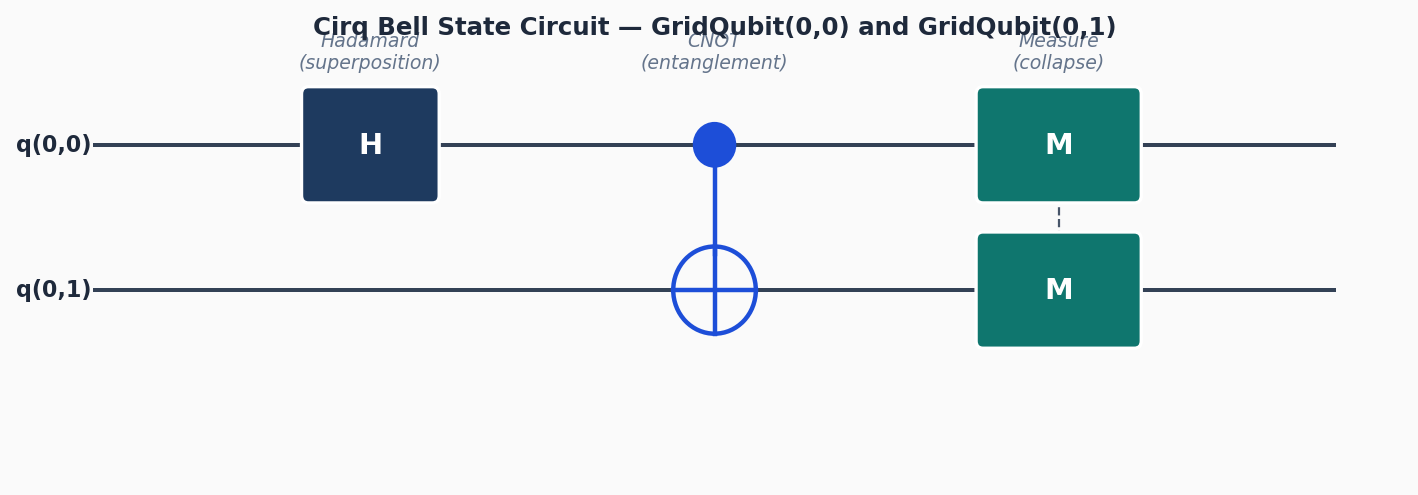
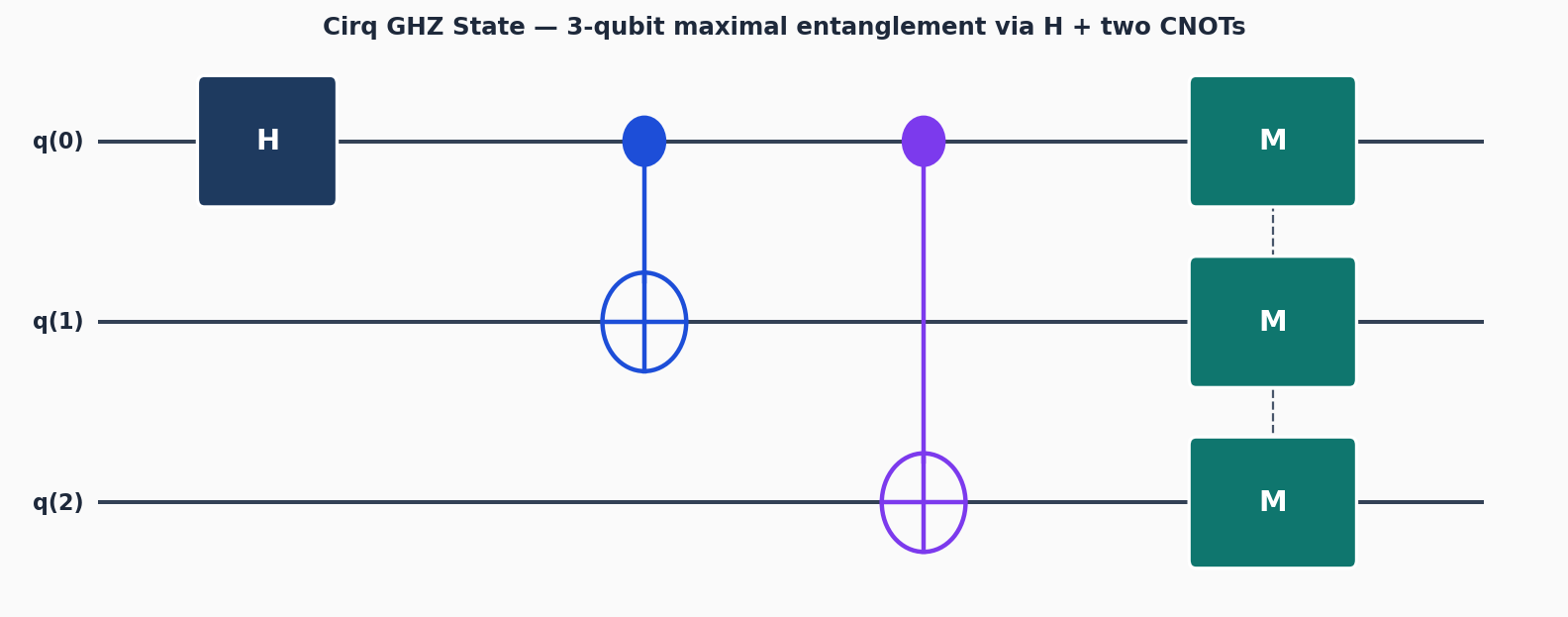
Why Cirq for VQE?
Cirq, developed at Google, gives you low-level control over quantum circuits that higher-level frameworks abstract away. For VQE research, this matters: you control exactly which gates are used, how parameters are bound, how circuits are swept, and how expectation values are computed. Cirq’s Sweep API makes parameter scans natural, and its tight integration with OpenFermion via the openfermion-cirq package gives you access to chemistry-motivated ansatze without sacrificing circuit-level transparency.
This tutorial builds VQE from the ground up: OpenFermion generates the H2 Hamiltonian, Cirq constructs and evaluates the parametric circuit, and scipy handles the classical optimization. We then implement the parameter shift rule for exact gradients and compare convergence with a gradient-free approach.
Installing Dependencies
pip install cirq openfermion openfermion-cirq scipy numpy
The openfermion-cirq package (also importable as cirq_google in some distributions; check ofermion.cirq for the canonical import path) provides UCCSD circuit generation compatible with Cirq’s gate model.
import cirq
import openfermion as of
import numpy as np
from scipy.optimize import minimize
Building the H2 Hamiltonian with OpenFermion
We use the same hardcoded H2 integrals from the STO-3G basis at bond length 0.74 angstrom:
# H2 molecular Hamiltonian, STO-3G, R=0.74A
# Build from InteractionOperator (1- and 2-body integrals)
one_body = np.array([[-1.2563, 0.0], [0.0, -0.4718]])
two_body = np.zeros((2, 2, 2, 2))
two_body[0, 0, 0, 0] = 0.6757
two_body[1, 1, 1, 1] = 0.6986
two_body[0, 0, 1, 1] = 0.6645
two_body[1, 1, 0, 0] = 0.6645
two_body[0, 1, 1, 0] = 0.1809
two_body[1, 0, 0, 1] = 0.1809
molecular_hamiltonian = of.InteractionOperator(
constant=0.7151,
one_body_tensor=one_body,
two_body_tensor=0.5 * two_body
)
fermionic_hamiltonian = of.get_fermion_operator(molecular_hamiltonian)
qubit_hamiltonian = of.jordan_wigner(fermionic_hamiltonian)
print(f"Number of Pauli terms: {len(qubit_hamiltonian.terms)}")
print("\nH2 qubit Hamiltonian:")
for term, coeff in sorted(qubit_hamiltonian.terms.items(), key=lambda x: -abs(x[1])):
if abs(coeff) > 1e-6:
label = " ".join(f"{op}{idx}" for idx, op in term) if term else "I"
print(f" {coeff.real:+.4f} {label}")
Defining the Qubits and Parametric Ansatz
The InteractionOperator built from these 2 spatial orbitals produces a Jordan-Wigner Hamiltonian acting on 2 qubits (one per spin-orbital: orbital 0 alpha and orbital 0 beta). The Hamiltonian is diagonal in the computational basis, and the ground state is the 2-electron state |11> where both spin-orbitals are occupied.
Cirq identifies qubits by objects, most commonly LineQubit (a 1D chain) or GridQubit (a 2D grid). For this 2-qubit H2 problem, LineQubit is natural:
# 2 qubits: spin-orbital 0 (alpha) and spin-orbital 1 (beta)
qubits = cirq.LineQubit.range(2)
q0, q1 = qubits
The circuit builder takes a params array and returns a concrete cirq.Circuit with those values baked in. This keeps the interface simple: the energy function and optimizer both just pass a numpy array of floats.
def make_vqe_circuit(params):
"""
2-parameter ansatz for H2.
Starts from the Hartree-Fock reference state |11>
(both spin-orbitals occupied) and applies parametric rotations.
The Hamiltonian is diagonal in Z, so independent Ry rotations
on each qubit are sufficient to reach the ground state.
"""
theta0, theta1 = params
circuit = cirq.Circuit([
# Hartree-Fock reference: set both qubits to |1>
cirq.X(q0),
cirq.X(q1),
# Parametric rotations
cirq.ry(theta0)(q0),
cirq.ry(theta1)(q1),
])
return circuit
print(make_vqe_circuit([0.0, 0.0]))
Computing Expectation Values
To evaluate <psi(theta)|H|psi(theta)>, decompose H into Pauli terms and measure each term’s expectation value separately. Cirq’s Simulator can compute exact statevector-based expectation values using cirq.PauliSum.
def openfermion_to_cirq_paulisum(qubit_op, cirq_qubits):
"""
Convert OpenFermion QubitOperator to Cirq PauliSum.
Assumes qubit index i maps to cirq_qubits[i].
"""
pauli_map = {"X": cirq.X, "Y": cirq.Y, "Z": cirq.Z}
terms = []
for term, coeff in qubit_op.terms.items():
if abs(coeff) < 1e-10:
continue
if not term:
# Identity term
terms.append(coeff.real * cirq.PauliString({}))
else:
pauli_string = cirq.PauliString(
{cirq_qubits[idx]: pauli_map[op] for idx, op in term},
coefficient=coeff.real
)
terms.append(pauli_string)
return cirq.PauliSum.from_pauli_strings(terms)
cirq_hamiltonian = openfermion_to_cirq_paulisum(qubit_hamiltonian, qubits)
print(f"Cirq PauliSum has {len(cirq_hamiltonian)} terms")
Now define the function that evaluates the energy for a given set of parameters:
simulator = cirq.Simulator()
def energy(params):
"""Evaluate <H> for given [theta0, theta1]."""
circuit = make_vqe_circuit(params)
result = simulator.simulate(circuit)
state_vector = result.final_state_vector
# Compute expectation value of each Pauli term
total_energy = 0.0
for pauli_string in cirq_hamiltonian:
expectation = pauli_string.expectation_from_state_vector(
state_vector,
qubit_map={q: i for i, q in enumerate(qubits)}
)
total_energy += expectation.real
return total_energy
# Test at a sample point
test_energy = energy([0.1, 0.2])
print(f"Energy at theta=[0.1, 0.2]: {test_energy:.6f} Hartree")
The Parameter Shift Rule
For quantum circuits with gates of the form exp(-i theta/2 * P) where P is a Pauli operator, the exact gradient can be computed using the parameter shift rule:
d<H>/d theta_k = [<H>(theta_k + pi/2) - <H>(theta_k - pi/2)] / 2
This requires only 2 circuit evaluations per parameter, exactly as many as a finite-difference approximation, but gives the exact analytical gradient rather than an approximation subject to numerical cancellation errors.
def gradient(params, shift=np.pi / 2):
"""
Compute gradient using the parameter shift rule.
Returns an array of partial derivatives.
"""
grads = np.zeros(len(params))
for k in range(len(params)):
params_plus = params.copy()
params_minus = params.copy()
params_plus[k] += shift
params_minus[k] -= shift
grads[k] = (energy(params_plus) - energy(params_minus)) / 2
return grads
# Verify gradient at a test point
params_test = np.array([0.5, -0.3])
g = gradient(params_test)
print(f"Gradient at [0.5, -0.3]: {g}")
# Compare with finite difference
eps = 1e-5
fd_grad = np.zeros(2)
e0 = energy(params_test)
for k in range(2):
params_fd = params_test.copy()
params_fd[k] += eps
fd_grad[k] = (energy(params_fd) - e0) / eps
print(f"Finite difference gradient: {fd_grad}")
print(f"Max absolute difference: {np.max(np.abs(g - fd_grad)):.2e}")
The parameter shift gradient and the finite difference gradient should agree to within numerical precision, but the parameter shift rule uses a shift of pi/2 (a large perturbation) rather than a tiny epsilon, making it robust to floating-point cancellation.
Classical Optimization Loop with scipy
energy_history = []
def energy_with_logging(params):
e = energy(params)
energy_history.append(e)
return e
# Gradient-free: Nelder-Mead
print("=== Nelder-Mead (gradient-free) ===")
energy_history.clear()
result_nm = minimize(
energy_with_logging,
x0=[0.01, 0.01],
method="Nelder-Mead",
options={"maxiter": 500, "xatol": 1e-7, "fatol": 1e-7}
)
print(f"Energy: {result_nm.fun:.6f} Hartree")
print(f"Params: {result_nm.x}")
print(f"Evaluations: {result_nm.nfev}")
# Gradient-based: L-BFGS-B with parameter shift gradients
print("\n=== L-BFGS-B with parameter shift gradients ===")
energy_history.clear()
result_lbfgs = minimize(
energy_with_logging,
x0=[0.01, 0.01],
method="L-BFGS-B",
jac=gradient,
options={"maxiter": 200, "ftol": 1e-12, "gtol": 1e-8}
)
print(f"Energy: {result_lbfgs.fun:.6f} Hartree")
print(f"Params: {result_lbfgs.x}")
print(f"Evaluations: {result_lbfgs.nfev}")
L-BFGS-B with exact parameter shift gradients typically converges in far fewer evaluations than Nelder-Mead for smooth landscapes like the H2 potential.
Energy Landscape Scan
With make_vqe_circuit building a fresh circuit per evaluation, scanning the landscape is a straightforward list comprehension. Fix theta1 at its optimal value and sweep theta0 across a full period:
# Scan theta0 from 0 to 2*pi with theta1 fixed at the optimal value
theta1_opt = result_lbfgs.x[1]
theta0_values = np.linspace(0, 2 * np.pi, 40)
sweep_energies = [energy([t0, theta1_opt]) for t0 in theta0_values]
print("\nEnergy landscape (theta0 scan):")
print(f"{'theta0':>10} {'Energy':>12}")
for t0, e in zip(theta0_values[::5], sweep_energies[::5]):
bar = "*" * int((e - min(sweep_energies)) / (max(sweep_energies) - min(sweep_energies)) * 30)
print(f"{t0:10.3f} {e:12.6f} {bar}")
print(f"\nMinimum energy at theta0={theta0_values[np.argmin(sweep_energies)]:.3f}: {min(sweep_energies):.6f}")
Verifying Against Exact Diagonalization
# Compute exact ground state via direct diagonalization of the Hamiltonian matrix
hamiltonian_matrix = of.get_sparse_operator(qubit_hamiltonian).toarray()
eigenvalues = np.linalg.eigvalsh(hamiltonian_matrix.real)
exact_ground_state_energy = eigenvalues[0]
print(f"\n=== Results Summary ===")
print(f"Exact ground state energy: {exact_ground_state_energy:.6f} Hartree")
print(f"VQE (Nelder-Mead): {result_nm.fun:.6f} Hartree")
print(f"VQE (L-BFGS-B + param shift): {result_lbfgs.fun:.6f} Hartree")
print(f"Chemical accuracy threshold: 0.001600 Hartree")
print(f"Nelder-Mead error: {abs(result_nm.fun - exact_ground_state_energy):.2e} Hartree")
print(f"L-BFGS-B error: {abs(result_lbfgs.fun - exact_ground_state_energy):.2e} Hartree")
Both should reach chemical accuracy (error below 0.0016 Hartree) for this simple 2-parameter H2 system. The L-BFGS-B optimizer typically gets to 10^-8 Hartree accuracy with fewer than 50 energy evaluations.
Typical Convergence Behavior
For H2 with this 2-parameter ansatz, the energy landscape has a single global minimum; convergence from any reasonable starting point is straightforward. For more complex ansatze or larger molecules:
- Nelder-Mead converges in O(N) iterations for N parameters on smooth problems but stalls on noisy or high-dimensional landscapes
- L-BFGS-B with parameter shift converges superlinearly near the minimum but requires the landscape to be smooth; on real hardware with shot noise, the gradient estimates become unreliable
- COBYLA is more robust to shot noise than L-BFGS-B but slower to converge; preferred when running on real hardware
- SPSA uses only 2 energy evaluations per gradient step regardless of N, making it ideal for large ansatze but requiring careful tuning of learning rate schedules
For deeper ansatze (UCCSD on molecules beyond H2), the barren plateau problem becomes severe: the gradient norm shrinks exponentially with circuit depth and qubit count, making gradient-based optimization ineffective without careful parameter initialization or layered training strategies.
Next Steps
Extending this Cirq VQE to larger molecules requires:
- Molecular integrals from PySCF via
openfermionpyscf(install separately) - UCCSD or ADAPT-VQE ansatze from
openfermion.circuits - Symmetry reduction to cut qubit count: particle-number and spin symmetry can halve the active space
- Noise simulation: replace
cirq.Simulator()with a noise model to see how gate errors affect convergence
The Cirq + OpenFermion stack gives you the transparency to experiment with all of these: you see every gate, every parameter binding, and every measurement outcome, making it the right choice for quantum chemistry algorithm research.
Was this tutorial helpful?