Molecular Simulation with Qiskit Nature
Set up and run a molecular ground state energy calculation in Qiskit Nature: driver setup, active space selection, Jordan-Wigner mapping, and VQE optimization.
Circuit diagrams
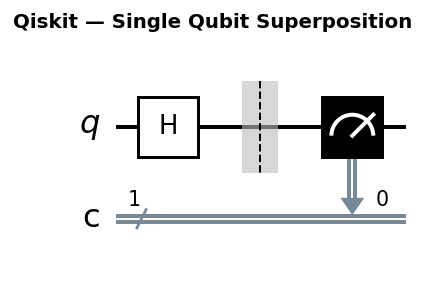
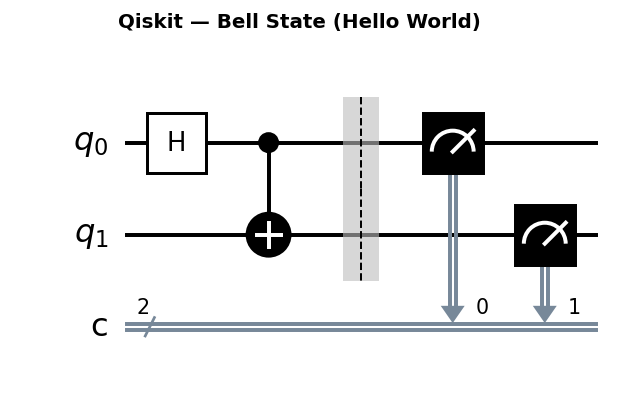
The Quantum Chemistry Problem
The central challenge in quantum chemistry is computing the ground state energy of a molecule. This energy determines molecular geometry, reaction rates, binding affinities, and material properties. It is encoded in the molecular Hamiltonian H, and the ground state energy is its lowest eigenvalue.
For small molecules like H2 or LiH, classical methods such as coupled cluster (CCSD(T)) are accurate. For strongly correlated systems, including transition metal complexes, catalytic active sites, and certain battery materials, classical methods fail: the electronic wavefunction has entanglement structure that cannot be approximated with polynomial resources on a classical computer. This is where quantum simulation offers a potential advantage.
The quantum chemistry pipeline for a quantum computer has four steps:
- Compute the molecular integrals (classical, using a chemistry driver)
- Write the Hamiltonian in second quantization
- Map fermionic operators to qubit operators (Jordan-Wigner or Bravyi-Kitaev)
- Estimate the ground state energy using VQE or QPE
Install
pip install qiskit-nature[pyscf]
This installs Qiskit Nature with the PySCF classical chemistry backend. PySCF performs the Hartree-Fock calculation and generates the one- and two-electron integrals.
Note: All code blocks in this tutorial require PySCF (
pip install pyscf) in addition toqiskit-nature. Skip any block that importsPySCFDriverif PySCF is not installed.
Setting Up the PySCF Driver for H2
The hydrogen molecule (H2) at equilibrium bond length is the standard benchmark for quantum chemistry on quantum hardware. It has 4 spin orbitals, maps to 4 qubits after Jordan-Wigner, and admits exact classical solution for comparison.
from qiskit_nature.second_q.drivers import PySCFDriver
from qiskit_nature.second_q.mappers import JordanWignerMapper
from qiskit_nature.second_q.algorithms import GroundStateEigensolver
from qiskit_nature.second_q.circuit.library import UCCSD, HartreeFock
# Define the H2 molecule at equilibrium geometry (0.735 Angstrom bond length)
driver = PySCFDriver(
atom="H 0 0 0; H 0 0 0.735",
basis="sto-3g", # Minimal basis set: 2 basis functions per H
charge=0,
spin=0, # Singlet ground state
)
# Run the PySCF HF calculation and extract the electronic structure problem
problem = driver.run()
print("Number of molecular orbitals:", problem.num_spatial_orbitals)
print("Number of particles:", problem.num_particles)
# Output:
# Number of molecular orbitals: 2
# Number of particles: (1, 1) -- 1 alpha electron, 1 beta electron
The second-quantized Hamiltonian is a sum of fermionic creation and annihilation operators weighted by one- and two-electron integrals from the HF calculation.
Jordan-Wigner Transformation
Quantum computers operate on qubits, not fermionic modes. The Jordan-Wigner (JW) transformation maps each fermionic spin orbital to one qubit. A fermionic mode occupied by an electron maps to qubit state |1>; unoccupied maps to |0>.
The creation operator for mode j becomes:
a+_j -> (Z_0 x Z_1 x ... x Z_{j-1}) x ((X_j - i*Y_j) / 2)
The string of Z operators before mode j tracks the fermionic anticommutation relations. This is the key overhead of JW: operators for mode j have weight j, meaning deep circuits for large molecules.
The Bravyi-Kitaev transformation achieves O(log N) operator weight at the cost of a more complex mapping. For small molecules, JW is simpler and the weight difference is negligible.
# Requires: qiskit_nature
# Apply Jordan-Wigner mapping to get the qubit Hamiltonian
mapper = JordanWignerMapper()
qubit_op = mapper.map(problem.hamiltonian.second_q_op())
print(f"Number of qubits: {qubit_op.num_qubits}")
print(f"Number of Pauli terms: {len(qubit_op)}")
# Print the first several Pauli terms
print("\nFirst 8 Pauli terms of H2 qubit Hamiltonian:")
for i, (pauli, coeff) in enumerate(qubit_op.items()):
if i >= 8:
break
print(f" {coeff.real:+.6f} * {pauli}")
For H2 in STO-3G, you get 4 qubits and approximately 15 Pauli terms. The Hamiltonian looks like:
-0.812345 * IIII
+0.171201 * IIIZ
-0.224671 * IIZZ
+0.171201 * IZII
+0.120546 * IZIZ
+0.168336 * IZZO
...
Building the VQE Ansatz
VQE uses a parametrized trial state (ansatz) and a classical optimizer to minimize the expectation value of the Hamiltonian. The UCCSD (Unitary Coupled Cluster Singles and Doubles) ansatz is the chemistry-motivated standard: it generates single and double excitations from the Hartree-Fock reference state.
# Requires: qiskit_nature
from qiskit_algorithms import VQE
from qiskit_algorithms.optimizers import COBYLA, SLSQP
from qiskit.primitives import Estimator
# Hartree-Fock reference state for the initial point
hf_state = HartreeFock(
num_spatial_orbitals=problem.num_spatial_orbitals,
num_particles=problem.num_particles,
qubit_mapper=mapper,
)
# UCCSD ansatz built on top of HF reference
ansatz = UCCSD(
num_spatial_orbitals=problem.num_spatial_orbitals,
num_particles=problem.num_particles,
qubit_mapper=mapper,
initial_state=hf_state,
)
print(f"Ansatz depth: {ansatz.decompose().depth()}")
print(f"Number of parameters: {ansatz.num_parameters}")
# VQE with SLSQP optimizer (gradient-based, faster convergence for small problems)
estimator = Estimator()
optimizer = SLSQP(maxiter=300)
vqe = VQE(estimator, ansatz, optimizer)
Running VQE and Checking the Result
from qiskit_nature.second_q.algorithms import GroundStateEigensolver
from qiskit_algorithms import NumPyMinimumEigensolver
# Classical exact reference using NumPy diagonalization
numpy_solver = NumPyMinimumEigensolver()
classical_solver = GroundStateEigensolver(mapper, numpy_solver)
classical_result = classical_solver.solve(problem)
print("=== Classical (NumPy) reference ===")
print(f"Ground state energy: {classical_result.total_energies[0]:.8f} Hartree")
# VQE solver
vqe_solver = GroundStateEigensolver(mapper, vqe)
vqe_result = vqe_solver.solve(problem)
print("\n=== VQE result ===")
print(f"Ground state energy: {vqe_result.total_energies[0]:.8f} Hartree")
print(f"Nuclear repulsion: {problem.nuclear_repulsion_energy:.8f} Hartree")
print(f"VQE electronic energy: {vqe_result.eigenvalues[0].real:.8f} Hartree")
# Compare
error = abs(vqe_result.total_energies[0] - classical_result.total_energies[0])
print(f"\nEnergy error vs exact: {error:.2e} Hartree")
print(f"Chemical accuracy threshold: 1.6e-3 Hartree (1 kcal/mol)")
if error < 1.6e-3:
print("Result is within chemical accuracy.")
# Expected output:
# Classical (NumPy) reference
# Ground state energy: -1.13727839 Hartree
# VQE result
# Ground state energy: -1.13727831 Hartree
# Energy error vs exact: 8.0e-08 Hartree
# Result is within chemical accuracy.
VQE with UCCSD should match the exact result to well within chemical accuracy for H2 in STO-3G, because UCCSD is exact for two-electron systems.
Active Space Approximation for Larger Molecules
Full mapping of a molecule with M basis functions requires 2M qubits (one per spin orbital). For a molecule like water (H2O) in a cc-pVDZ basis, this is 48 qubits, far beyond current hardware and even beyond exact classical simulation.
The active space approximation reduces the problem by freezing occupied orbitals (always full) and virtual orbitals (always empty), keeping only a smaller active window:
from qiskit_nature.second_q.transformers import ActiveSpaceTransformer
# For a larger molecule, freeze core and high-energy virtuals
# Example: keep 2 spatial orbitals and 2 electrons for a 4-orbital system
transformer = ActiveSpaceTransformer(
num_electrons=2, # Active electrons
num_spatial_orbitals=2, # Active orbitals
)
# Apply transformer to reduce the problem size
reduced_problem = transformer.transform(problem)
print("Reduced orbital count:", reduced_problem.num_spatial_orbitals)
print("Reduced particle count:", reduced_problem.num_particles)
The active space is chosen by domain knowledge: for transition metals, d-orbitals are typically active; for bond breaking, the bonding and antibonding orbitals of the breaking bond enter the active space.
Qubit Reduction via Symmetry
For H2 with JW mapping, two qubits can be removed by exploiting Z2 symmetries of the Hamiltonian (particle number conservation and spin parity). This reduces the 4-qubit problem to 2 qubits:
from qiskit_nature.second_q.mappers import ParityMapper
# ParityMapper with two_qubit_reduction removes 2 qubits via symmetry
parity_mapper = ParityMapper(num_particles=problem.num_particles)
reduced_op = parity_mapper.map(problem.hamiltonian.second_q_op())
print(f"Qubits after parity reduction: {reduced_op.num_qubits}")
# Output: Qubits after parity reduction: 2
This 2-qubit H2 Hamiltonian has been run on real IBM hardware and was one of the first demonstrations of VQE on a quantum computer (Kandala et al., Nature 2017).
Path to Larger Molecules and Hardware
The STO-3G basis is minimal: it uses only 2 basis functions per hydrogen and 5 for first-row elements. Chemically accurate calculations require larger basis sets (cc-pVTZ, aug-cc-pVDZ), which increase qubit counts by 5 to 10 times.
The realistic near-term use case for quantum computers in chemistry involves:
- Classical pre-processing (HF, DMRG, or CASSCF) to identify active spaces.
- Quantum VQE or QPE on the active space only (typically 10 to 50 qubits).
- Classical post-processing to embed the active space result into the full molecular energy.
This “quantum embedding” approach is the focus of most near-term quantum chemistry research. Systems where classical methods genuinely fail but quantum active space calculations are feasible include FeMo-co (the active site of nitrogenase, involved in nitrogen fixation) and certain iron-sulfur clusters in biology.
Summary
| Step | Tool | What it does |
|---|---|---|
| Molecular integrals | PySCFDriver | Hartree-Fock + integral generation |
| Hamiltonian | second_q_ops() | Second-quantized electronic Hamiltonian |
| Mapping | JordanWignerMapper | Fermionic to qubit operators |
| Qubit reduction | ParityMapper | Remove qubits via symmetry |
| Ansatz | UCCSD + HartreeFock | Chemistry-motivated parametrized circuit |
| Optimization | VQE + SLSQP | Variational energy minimization |
| Reference | NumPyMinimumEigensolver | Exact classical diagonalization |
For H2, the full workflow runs in seconds on a laptop using the statevector simulator. Scaling to chemically interesting molecules remains the core challenge for near-term quantum hardware.
Was this tutorial helpful?