Molecular Simulation with OpenFermion and Qiskit
Build a complete quantum chemistry pipeline: encode the H2 Hamiltonian with OpenFermion, transform it to qubits via Jordan-Wigner, then find the ground state energy using VQE on Qiskit's statevector simulator.
Circuit diagrams
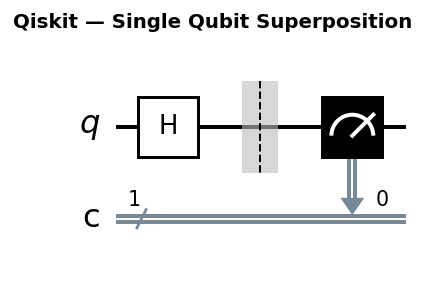
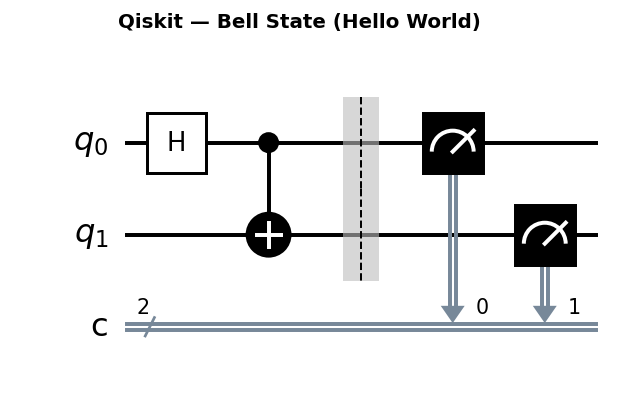
Why Quantum Chemistry is a Natural Quantum Computing Application
Simulating molecules on a classical computer is exponentially hard. The exact wavefunction of a molecule with N electrons lives in a Hilbert space of dimension 2^N; you need to store and manipulate exponentially large vectors. Methods like full configuration interaction (FCI) scale as O(N! / (k!(N-k)!)) in the number of basis functions, quickly becoming intractable beyond a dozen electrons.
Quantum computers offer a natural encoding: map each molecular orbital to a qubit. The quantum state of N qubits has 2^N amplitudes, exactly the right representation for N electrons in N orbitals. Quantum simulation of chemistry was, in fact, the original motivation Feynman had in mind when he proposed quantum computing.
Second Quantization: The Language of Quantum Chemistry
Classical quantum chemistry uses the wavefunction picture: track every electron explicitly. Second quantization reformulates this in terms of creation and annihilation operators acting on a vacuum state.
A creation operator a†_p adds an electron to orbital p. An annihilation operator a_p removes one. The molecular Hamiltonian in second quantization is:
H = sum_{pq} h_{pq} a†_p a_q + 0.5 * sum_{pqrs} h_{pqrs} a†_p a†_q a_r a_s
where h_{pq} are one-electron integrals (kinetic energy + nuclear attraction) and h_{pqrs} are two-electron repulsion integrals. These integrals come from classical quantum chemistry packages like PySCF or PSI4.
The advantage: the Hamiltonian is now expressed in a fixed-size algebraic form regardless of how many electrons are present. The complication: fermionic operators obey anticommutation relations, not the commutation relations natural to qubits.
The Jordan-Wigner Transformation
To run quantum chemistry on a qubit processor, you must map fermionic operators to qubit operators. The Jordan-Wigner (JW) transformation is the most common choice:
a_j = (tensor of Z's from qubit 0 to j-1) x (X_j + i Y_j) / 2
Each fermionic mode maps to one qubit. The occupation of orbital j is encoded in the |0> or |1> state of qubit j. The string of Z operators to the left of position j enforces the fermionic anticommutation statistics; this is the price of embedding fermionic algebra into spin algebra.
The result is a sum of Pauli strings: terms like X0 Z1 Y2 I3, each with a real coefficient. This Pauli sum form is exactly what VQE needs to compute expectation values.
Installing the Tools
pip install openfermion qiskit qiskit-aer qiskit-algorithms scipy
For computing molecular integrals from scratch you would also install pyscf and openfermionpyscf, but for this tutorial we hardcode the H2 Hamiltonian integrals at a fixed bond length to keep dependencies minimal.
Building the H2 Hamiltonian
Hydrogen molecule (H2) is the canonical benchmark. At a bond length of 0.74 angstroms, the STO-3G basis gives a 2-spatial-orbital (2-qubit after JW) Hamiltonian when using spatial-orbital integrals. The integrals below are for the two spatial orbitals (bonding and antibonding MOs); spin degrees of freedom are folded in via the two-body tensor.
import openfermion as of
import numpy as np
# H2 molecular integrals at R=0.74 angstrom, STO-3G basis
# (precomputed via PySCF / openfermion-pyscf)
n_orbitals = 2
n_electrons = 2
# One-body integrals (alpha spin)
one_body_coefficients = np.array([
[-1.2563, 0.0],
[ 0.0, -0.4718]
])
# Two-body integrals (physicists' notation: <pq|rs>)
two_body_coefficients = np.zeros((2, 2, 2, 2))
two_body_coefficients[0, 0, 0, 0] = 0.6757
two_body_coefficients[1, 1, 1, 1] = 0.6986
two_body_coefficients[0, 0, 1, 1] = 0.6645
two_body_coefficients[1, 1, 0, 0] = 0.6645
two_body_coefficients[0, 1, 1, 0] = 0.1809
two_body_coefficients[1, 0, 0, 1] = 0.1809
# Build the FermionOperator Hamiltonian
molecular_hamiltonian = of.InteractionOperator(
constant=0.7151,
one_body_tensor=one_body_coefficients,
two_body_tensor=0.5 * two_body_coefficients
)
fermionic_hamiltonian = of.get_fermion_operator(molecular_hamiltonian)
print("Fermionic Hamiltonian terms (first 5):")
for term, coeff in list(fermionic_hamiltonian.terms.items())[:5]:
print(f" {coeff:.4f} {term}")
The FermionOperator represents the Hamiltonian as a sum of products of creation and annihilation operators. The constant 0.7151 Hartree is the nuclear repulsion energy for H2 at 0.74 angstroms.
Applying the Jordan-Wigner Transformation
# Requires: openfermion
# Transform to qubit operators
qubit_hamiltonian = of.jordan_wigner(fermionic_hamiltonian)
print(f"\nNumber of Pauli terms after JW: {len(qubit_hamiltonian.terms)}")
print("\nQubit Hamiltonian (first 8 terms):")
for term, coeff in list(qubit_hamiltonian.terms.items())[:8]:
if abs(coeff) > 1e-6:
pauli_string = " ".join(f"{op}_{idx}" for idx, op in term) if term else "I"
print(f" {coeff.real:+.4f} {pauli_string}")
H2 in STO-3G with spatial-orbital integrals produces 4 Pauli terms after JW transformation acting on 2 qubits. The terms are diagonal (all Z or identity) because the spatial-orbital Hamiltonian has no off-diagonal hopping at this level of representation. A full spin-orbital treatment would require 4 qubits and ~15 Pauli terms including X and Y contributions from the exchange integrals.
Converting to Qiskit’s SparsePauliOp
Qiskit uses SparsePauliOp internally. We convert from OpenFermion’s format:
# Requires: openfermion
from qiskit.quantum_info import SparsePauliOp
def openfermion_to_qiskit(qubit_op, n_qubits):
"""Convert OpenFermion QubitOperator to Qiskit SparsePauliOp."""
pauli_list = []
for term, coeff in qubit_op.terms.items():
if abs(coeff) < 1e-10:
continue
# Build the Pauli string (Qiskit uses reversed qubit ordering)
pauli_chars = ["I"] * n_qubits
for qubit_idx, pauli_char in term:
pauli_chars[n_qubits - 1 - qubit_idx] = pauli_char
pauli_str = "".join(pauli_chars)
pauli_list.append((pauli_str, coeff.real))
return SparsePauliOp.from_list(pauli_list)
n_qubits = of.count_qubits(qubit_hamiltonian)
qiskit_hamiltonian = openfermion_to_qiskit(qubit_hamiltonian, n_qubits)
print(f"\nConverted to SparsePauliOp with {len(qiskit_hamiltonian)} terms")
print(qiskit_hamiltonian)
Note the qubit ordering reversal: OpenFermion indexes qubits from left to right, while Qiskit uses the convention where qubit 0 is the rightmost character in a Pauli string.
Designing the Ansatz
For H2 with 2 spatial orbitals mapped to 2 qubits, a two-parameter hardware-efficient ansatz that spans the relevant subspace is:
from qiskit.circuit import QuantumCircuit, ParameterVector
def h2_ansatz():
"""
Two-parameter ansatz for H2 ground state (2-qubit, spatial-orbital basis).
Qubit 0 represents the bonding MO; qubit 1 the antibonding MO.
"""
theta = ParameterVector("theta", 2)
qc = QuantumCircuit(2)
# Prepare reference state with one electron in the bonding orbital
qc.x(0)
# Parametric rotations to explore superposition
qc.ry(theta[0], 0)
qc.ry(theta[1], 1)
# Entanglement layer
qc.cx(0, 1)
return qc
ansatz = h2_ansatz()
print(ansatz.draw("text"))
print(f"\nNumber of parameters: {ansatz.num_parameters}")
Running VQE with the Estimator Primitive
# Requires: openfermion
from qiskit_algorithms import VQE
from qiskit_algorithms.optimizers import COBYLA, SLSQP
from qiskit.primitives import StatevectorEstimator
# Statevector estimator gives exact expectation values (no shot noise)
estimator = StatevectorEstimator()
optimizer = COBYLA(maxiter=500, rhobeg=0.5)
vqe = VQE(
estimator=estimator,
ansatz=ansatz,
optimizer=optimizer,
initial_point=[0.0, 0.0]
)
result = vqe.compute_minimum_eigenvalue(qiskit_hamiltonian)
print(f"\nVQE ground state energy: {result.eigenvalue:.6f} Hartree")
print(f"Optimal parameters: {result.optimal_parameters}")
print(f"Function evaluations: {result.cost_function_evals}")
Comparing with the Exact Solution
Verify against the exact diagonalization (FCI) result:
# Requires: openfermion
from qiskit_algorithms import NumPyMinimumEigensolver
exact_solver = NumPyMinimumEigensolver()
exact_result = exact_solver.compute_minimum_eigenvalue(qiskit_hamiltonian)
fci_energy = exact_result.eigenvalue.real
vqe_energy = result.eigenvalue.real
chemical_accuracy = 0.0016 # Hartree (1 kcal/mol)
print(f"\nFCI (exact) energy: {fci_energy:.6f} Hartree")
print(f"VQE energy: {vqe_energy:.6f} Hartree")
print(f"Error: {abs(vqe_energy - fci_energy):.6f} Hartree")
print(f"Chemical accuracy: {chemical_accuracy:.4f} Hartree")
print(f"Within chemical accuracy: {abs(vqe_energy - fci_energy) < chemical_accuracy}")
For H2 with this 2-parameter ansatz, VQE should reach the FCI energy to within 10^-5 Hartree, well within the 1.6 millihartree chemical accuracy threshold.
What Comes Next
The H2 example is illustrative but trivially small. Practical quantum chemistry targets molecules where classical FCI is intractable: systems with 50-200 electrons in an active space of 20-50 orbitals. Reaching that regime requires:
- Better ansatze: UCCSD or ADAPT-VQE to reduce parameter count while staying close to the exact ground state
- Larger qubit counts: ~40 qubits for a 20-orbital active space
- Error mitigation: zero-noise extrapolation or probabilistic error cancellation to suppress hardware noise
- Symmetry reduction: exploiting particle number and spin symmetry to reduce the effective Hilbert space
OpenFermion integrates with PySCF via openfermion-pyscf to generate integrals from actual molecular geometries. Replace the hardcoded integrals above with a MolecularData object and a PySCF SCF calculation to simulate real molecules like LiH, BeH2, or small organic radicals.
Was this tutorial helpful?